Role of Cytokinin in Regulating Shoot Apical Meristem Function in Rice (NSF Plant Genome)
Joe Kieber (PI); Josh Strable, coPI (NCSU); Eric Schaller, coPI (Dartmouth)
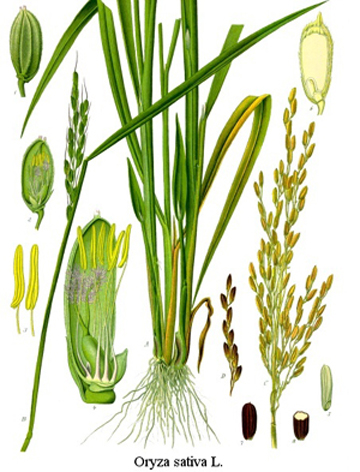
In contrast to Arabidopsis thaliana, which is a dicot (class Magnoliopsida), rice (Oryza sativa) is a monocot (class Liliopsida)(What's the difference?). Various lines of evidence suggest that the function of two-component elements may be distinct in these two plant classes (e.g. Giulini et al. 2004 Nature 430:1031-4; Burr et al, 2020 Development 147(20):dev191734). This project aims to elucidate the role of cytokinin in rice growth and development, focusing on cytokinin’s role in regulating apical meristem function. Using single cell approaches, we will determine the transcription modules regulated by cytokinin across the shoot apical/inflorescence meristem (SAM/IM).
Morphology of rice (Source:www.plant-pictures.de)
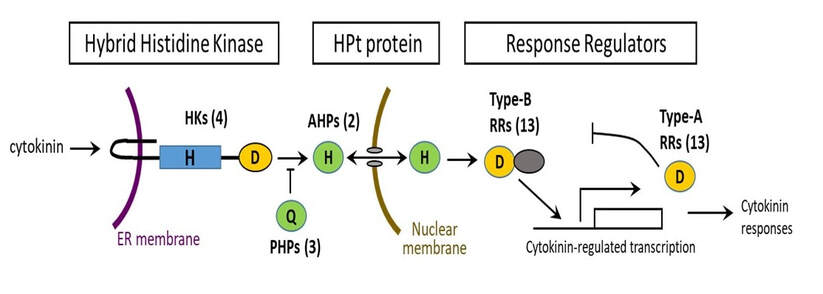
The rice cytokinin signaling pathway is similar to that of Arabidopsis. We have focused on the disruption of the OsAHPs (Fig. 3), which are positive elements represented by only two genes in the rice genome, and the type-A OsRRs, which, at least in Arabidopsis are negative regulatyors of cytokinin signaling.
Cytokinin signaling in rice. The numbers in parenthesis indicate the number of genes encoding each element present in the rice genome
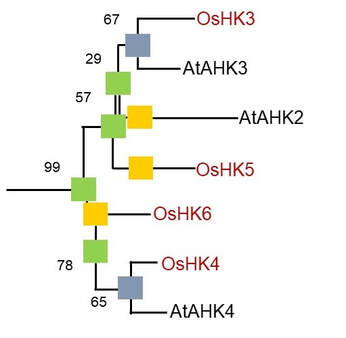
A phylogentic analysis of HK proteins from rice and Arabidopsis.
Project Objectives
We hypothesize that cytokinin modulates the function of apical meristems by altering specific sets of target genes across developmental time and space in concert with other signals. We will employ an integrated set of experiments to reveal cytokinin’s role in regulating apical meristem function and to elucidate the transcription modules regulated by cytokinin across the shoot apical/inflorescence meristem (SAM/IM).
(1) Generate foundational data of gene expression and chromatin states across the SAM and IM from wild-type rice using scRNA-seq and snATAC-seq.
(2) Determine how disruption of cytokinin perception alters rice SAM and IM development and gene expression profiles at single cell level.
(3) Determine the role of the cytokinin-regulated type-B RRs in rice SAM and IM development.
Broader impacts include developing and implementing local interactive programs that foster science education to the general public, which include working with museum, science center, and gallery and art center spaces.
Project Highlights
We hypothesize that cytokinin modulates the function of apical meristems by altering specific sets of target genes across developmental time and space in concert with other signals. We will employ an integrated set of experiments to reveal cytokinin’s role in regulating apical meristem function and to elucidate the transcription modules regulated by cytokinin across the shoot apical/inflorescence meristem (SAM/IM).
(1) Generate foundational data of gene expression and chromatin states across the SAM and IM from wild-type rice using scRNA-seq and snATAC-seq.
(2) Determine how disruption of cytokinin perception alters rice SAM and IM development and gene expression profiles at single cell level.
(3) Determine the role of the cytokinin-regulated type-B RRs in rice SAM and IM development.
Broader impacts include developing and implementing local interactive programs that foster science education to the general public, which include working with museum, science center, and gallery and art center spaces.
Project Highlights
- The CRISPR/Cas9 system is proving highly effective for generating loss-of-function mutations in rice. Using this, we have developed an extensive collection of cytokinin receptor mutants in rice.
- Analysis of hk mutants reveals a critical function for cytokinin in shoot apical meristem and inflorescence meristem formation and development.
Publications supported by this award:
Some manuscripts describing two-component elements in rice:
Tsai YC, Weir NR, Hill K, Zhang W, Kim HJ, Shiu SH, Schaller GE, Kieber JJ. (2012) Characterization of genes involved in cytokinin signaling and metabolism from rice. Plant Physiol. 158:1666-84
Du, L., Jiao, F., Chu, J., Jin, G., Chen, M., Wu, P. (2007) The two-component signal system in rice (Oryza sativa L.): a genome-wide study of cytokinin signal perception and transduction.
Genomics. 89:697-707
Schaller, G.E., Doi, K., Hwang, I., Kieber, J.J., Khurana, J.P., Kurata, N., Mizuno, T., Pareek, A., Shiu, S.H., Wu P., Yip, W.K. (2007) Nomenclature for two-component signaling elements of rice. Plant Physiol. 143:555-7.
Ito, Y. and Kurata, N. (2006) Identification and characterization of cytokinin-signalling gene families in rice. Gene 382: 57-65.
Pareek, A., Singh, A., Kumar, M., Kushwaha, H.R., Lynn, A.M., Singla-Pareek, S.L. (2006) Whole genome analysis of Oryza sativa l. reveals similar architecture of two-component-signaling-machinery with Arabidopsis. Plant Physiol.142(2):380-97.
Jain, M., Tyagi, A.K., Khurana, J.P. (2006) Molecular characterization and differential expression of cytokinin-responsive type-A response regulators in rice (Oryza sativa). BMC Plant Biol. 6:1
This project is supported by the National Science Foundation Plant Genome Research Program
Some manuscripts describing two-component elements in rice:
Tsai YC, Weir NR, Hill K, Zhang W, Kim HJ, Shiu SH, Schaller GE, Kieber JJ. (2012) Characterization of genes involved in cytokinin signaling and metabolism from rice. Plant Physiol. 158:1666-84
Du, L., Jiao, F., Chu, J., Jin, G., Chen, M., Wu, P. (2007) The two-component signal system in rice (Oryza sativa L.): a genome-wide study of cytokinin signal perception and transduction.
Genomics. 89:697-707
Schaller, G.E., Doi, K., Hwang, I., Kieber, J.J., Khurana, J.P., Kurata, N., Mizuno, T., Pareek, A., Shiu, S.H., Wu P., Yip, W.K. (2007) Nomenclature for two-component signaling elements of rice. Plant Physiol. 143:555-7.
Ito, Y. and Kurata, N. (2006) Identification and characterization of cytokinin-signalling gene families in rice. Gene 382: 57-65.
Pareek, A., Singh, A., Kumar, M., Kushwaha, H.R., Lynn, A.M., Singla-Pareek, S.L. (2006) Whole genome analysis of Oryza sativa l. reveals similar architecture of two-component-signaling-machinery with Arabidopsis. Plant Physiol.142(2):380-97.
Jain, M., Tyagi, A.K., Khurana, J.P. (2006) Molecular characterization and differential expression of cytokinin-responsive type-A response regulators in rice (Oryza sativa). BMC Plant Biol. 6:1
This project is supported by the National Science Foundation Plant Genome Research Program